Cardiosim-Tox: An Interpretable Multitask Deep Learning QSAR Platform with Multi-Modal Feature Fusion for hERG, Cav1.2, and Nav1.5 Blockade Risk and Potency Prediction
Cardiosim-Tox is an interpretable multitask deep learning QSAR platform for predicting cardiac ion-channel blockade risk and potency across three major cardiac ion channels: hERG, Cav1.2, and Nav1.5.
This repository contains the tutorial notebooks used for molecular descriptor calculation, chemical diversity analysis, applicability domain analysis, model training, and model evaluation for binary classification and regression tasks.
Drug-induced cardiotoxicity remains one of the major causes of drug attrition and post-market withdrawal. A key mechanism underlying this toxicity is the blockade of cardiac ion channels, particularly hERG, Cav1.2, and Nav1.5, which are closely associated with cardiac electrophysiological disturbances.
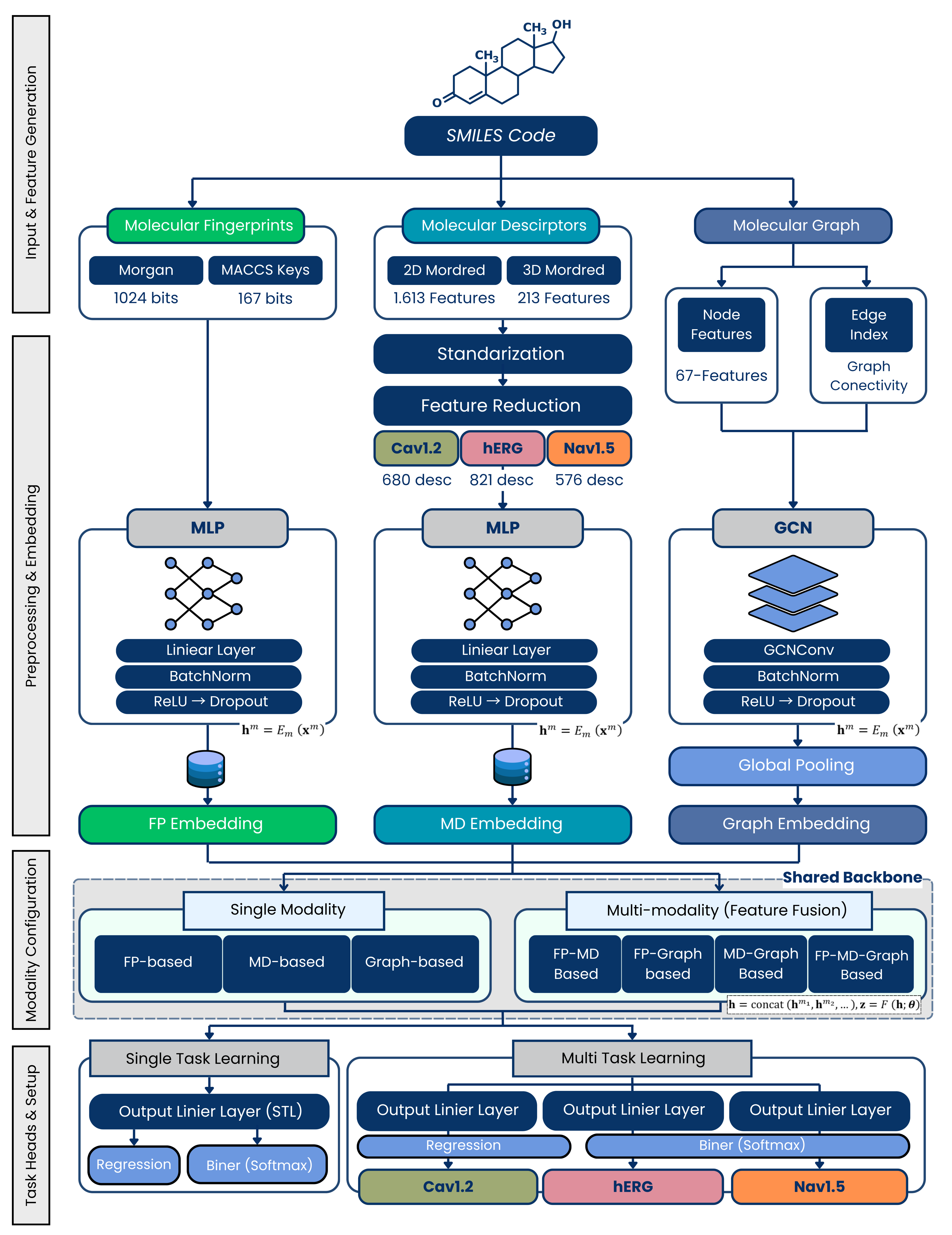
Overall framework of the Cardiosim-Tox platform for multi-modal feature fusion, multitask learning, and prediction of cardiac ion-channel blockade risk and potency.
To support early-stage cardiotoxicity screening, this study develops Cardiosim-Tox, a multi-modal and multitask QSAR platform that predicts:
-
Blockade risk
Binary classification of compounds as blockers or non-blockers. -
Blockade potency
Regression prediction of pIC₅₀ values.
The platform integrates multiple molecular representations, including:
- Molecular fingerprints
- Mordred molecular descriptors
- Graph-based molecular encodings
The model is designed under a multitask learning framework, allowing shared learning across multiple cardiac ion-channel endpoints. This enables the model to capture common molecular patterns associated with ion-channel blockade while maintaining endpoint-specific prediction capacity.
- Multitask deep learning for hERG, Cav1.2, and Nav1.5 prediction
- Binary classification for ion-channel blockade risk
- Regression modeling for pIC₅₀ potency prediction
- Multi-modal molecular feature fusion
- Molecular descriptor calculation workflow
- Chemical diversity analysis using UMAP
- Applicability domain analysis
- External validation and performance evaluation
- Interpretable modeling framework supporting QSAR-based cardiotoxicity screening
This repository includes the following tutorial notebooks:
| Notebook | Description |
|---|---|
Biner_Descriptor Calculation.ipynb |
Calculates molecular descriptors and prepares features for binary classification tasks. |
Chemical Diversity Test_UMAP.ipynb |
Performs chemical space visualization and diversity analysis using UMAP. |
Training_MTL_Binary Classification.ipynb |
Trains multitask learning models for binary ion-channel blockade classification. |
Evaluation_MTL_Binary Classification.ipynb |
Evaluates trained multitask classification models. |
Training_MTL_Regression.ipynb |
Trains multitask learning models for pIC₅₀ regression prediction. |
Evaluation_MTL_Regression.ipynb |
Evaluates trained multitask regression models. |
Aplicability Domain_Analysis.ipynb |
Performs applicability domain analysis for model reliability assessment. |
Note: The notebook name
Aplicability Domain_Analysis.ipynbfollows the current file name in this repository. For consistency, it may be renamed toApplicability Domain_Analysis.ipynb.
The development and testing datasets used in this study are available on Zenodo:
Please download the dataset from the Zenodo record before running the notebooks. After downloading, place the dataset files in the appropriate data directory used by each notebook, or update the file paths inside the notebooks according to your local folder structure.
The dataset includes data used for model development and testing of the Cardiosim-Tox platform for hERG, Cav1.2, and Nav1.5 blockade risk and potency prediction.
The notebooks should be run in the following order:
Run:
Biner_Descriptor Calculation.ipynb
This notebook prepares molecular descriptors and molecular features used for model development. It may include preprocessing of SMILES strings, descriptor calculation, feature cleaning, and dataset preparation.
Run:
Chemical Diversity Test_UMAP.ipynb
This notebook analyzes the chemical diversity of the dataset and visualizes the molecular chemical space using UMAP. This step helps evaluate whether training, validation, and external test compounds occupy similar or distinct regions of chemical space.
Run:
Training_MTL_Binary Classification.ipynb
This notebook trains the multitask deep learning model for binary classification of ion-channel blockade risk across hERG, Cav1.2, and Nav1.5.
The output of this notebook may include trained model files, training history, loss curves, and classification-related outputs.
Run:
Evaluation_MTL_Binary Classification.ipynb
This notebook evaluates the trained binary classification model using metrics such as:
- Accuracy
- Sensitivity
- Specificity
- Precision
- F1-score
- ROC-AUC
- PR-AUC
The evaluation is performed for each cardiac ion-channel endpoint.
Run:
Training_MTL_Regression.ipynb
This notebook trains the multitask deep learning model for potency prediction using pIC₅₀ values.
The model learns regression tasks for cardiac ion-channel blockade potency prediction.
Run:
Evaluation_MTL_Regression.ipynb
This notebook evaluates regression model performance using metrics such as:
- R²
- RMSE
- MAE
Run:
Aplicability Domain_Analysis.ipynb
This notebook evaluates whether query compounds fall within the applicability domain of the trained QSAR models. Applicability domain analysis is important for assessing the reliability of model predictions, especially for external or structurally novel compounds.
git clone https://github.com/YOUR_USERNAME/Cardiosim-Tox.git
cd Cardiosim-ToxReplace YOUR_USERNAME with your GitHub username or organization name.
Using conda:
conda create -n cardiosimtox python=3.10
conda activate cardiosimtoxOr using venv:
python -m venv cardiosimtox_env
cardiosimtox_env\Scripts\activateFor macOS/Linux:
source cardiosimtox_env/bin/activateInstall the main dependencies:
pip install numpy pandas scipy scikit-learn matplotlib seaborn tqdm jupyter notebook
pip install rdkit mordred
pip install torch torchvision torchaudio
pip install torch-geometric
pip install shap umap-learnDepending on your system and CUDA version, PyTorch and PyTorch Geometric installation may require specific commands. Please refer to the official installation pages if GPU support is needed.
jupyter notebookThen open and run the notebooks according to the recommended workflow.
The notebooks require curated molecular datasets containing molecular structures and endpoint labels or pIC₅₀ values.
Typical input columns may include:
| Column | Description |
|---|---|
SMILES |
Molecular structure in SMILES format |
hERG |
Binary label or pIC₅₀ value for hERG |
Cav1.2 |
Binary label or pIC₅₀ value for Cav1.2 |
Nav1.5 |
Binary label or pIC₅₀ value for Nav1.5 |
The exact column names should be adjusted according to the dataset format used in each notebook.
Depending on the notebook, the workflow may generate:
- Processed descriptor files
- Molecular fingerprint matrices
- Training and validation datasets
- Trained deep learning model files
- Model performance tables
- ROC and PR curves
- Regression scatter plots
- UMAP chemical space visualizations
- Applicability domain results
- Prediction outputs for external compounds
Cardiosim-Tox supports two main prediction tasks.
The binary classification task predicts whether a compound is likely to block a given cardiac ion channel.
Endpoints:
- hERG blocker / non-blocker
- Cav1.2 blocker / non-blocker
- Nav1.5 blocker / non-blocker
The regression task predicts ion-channel blockade potency as pIC₅₀.
Endpoints:
- hERG pIC₅₀
- Cav1.2 pIC₅₀
- Nav1.5 pIC₅₀
Applicability domain analysis is included to assess whether a compound is located within the chemical space covered by the training data.
Predictions for compounds outside the applicability domain should be interpreted carefully, as model reliability may decrease for structurally distant molecules.
If you use this repository, model, or workflow in your research, please cite:
Nursyafi FS, Kang B, Eom J, Izza RN, Fauzan AL, Putri NH, Hanum UL, Olivia Y, Rose SA, Lim KM.
Cardiosim-Tox: An Interpretable Multitask Deep Learning QSAR Platform with Multi-Modal Feature Fusion for hERG, Cav1.2, and Nav1.5 Blockade Risk and Potency Prediction.
Once the article is published, please update this section with the final journal citation and DOI.
Fauzan Syarif Nursyafi
Kang-Byunggyu
Eom-Junhyeok
Rahmafatin Nurul Izza
Abdul Latif Fauzan
Nadilla Hafani Putri
Ulfa Latifa Hanum
Yossi Olivia
Sanjida Afrin Rose
Ki Moo Lim
Ki Moo Lim
Department of Biomedical Engineering
Kumoh National Institute of Technology
Gumi, Republic of Korea
Email: kmlim@kumoh.ac.kr
Cardiosim-Tox is intended for research purposes only. Model predictions should not be used as the sole basis for clinical, regulatory, or safety-critical decisions. Experimental validation is recommended for compounds of interest.
cardiotoxicity, QSAR, multitask learning, deep learning, hERG, Cav1.2, Nav1.5, ion-channel blockade, molecular descriptors, molecular fingerprints, graph neural network, applicability domain, computational toxicology